Integrated Global Health Surveillance and Response through Multi-Source Technologies
Paul O. Kwon
Introduction
Since 1980, one to three new human infections have been identified annually. These Emerging Infectious Diseases (EIDs) impact social, political, economic and environmental arenas. In response, the International Health Regulations (2005) provide a legally binding framework to guide international member states of the World Health Organization (WHO) to “detect, assess, and report” potential outbreaks and other public health emergencies. More importantly, infectious disease is the one of the leading cause of morbidity and mortality around the world impacting socioeconomic stability. Globally, the Avian flu pandemic ($3.78 trillion) and Ebola outbreak ($32.6 billion) revealed critical shortcomings in mobilizing and leveraging public and private sectors that threaten global public health and security. More recently, vector borne diseases such as Chikungunya and Zika virus have threatened migration towards the southern borders of North America. Thus, forecasting outbreaks is crucial in planning for an effective response strategy.
Similarly, the U.S. has also developed key policy milestones for greater interagency support for national and global EID efforts (1997 Presidential Directive; 2014 Global Health Security Agenda). Interestingly, a common virus such as the seasonal flu has a significant annual impact within the U.S. population (up to 5-20%) and healthcare (over 200,000 hospitalizations and 3,000-49,000 deaths with over $87 billion in reduced economic output). Understanding the dynamics of disease forecasting is therefore essential for prevention and mitigation strategies; however, there must be a mobilization towards a multidisciplinary and holistic approach in these proactive measures. In the face of budget constraints and limited resourcing, truly there is a need for an Integrated Global Health Surveillance and Response Program through a complex web of surveillance, education, and interactive coordination (“network of networks”).
Background
In the past decade, there has been a significant technologic advancement in disease surveillance capabilities. Several bioinformatics science and technology enterprises have developed a number of tools that leverage data science and existing open source data. With the ubiquitous desire to seek technologic advantages (internet, social media, smart phone and apps), the social network provides a wealth of information that can enhance the Warfighters’ force health protection and offer solutions towards predictable models in reference to EIDs and environmental threats to major populations.
Worldwide, six billion persons have mobile phones according to a recent U.N. report and there has been a rapid global proliferation of cellular phones in the last three decades. According to the PEW Research, more than 90% of all U.S. adults own a cell phone and 50% download apps. Trends also include a rise in smart phone purchasing (up to 1.45 billion in 2015). Consumers are spending more time on their apps than ever before, the abundance of mobile devices has transformed the information age, and the Neilson Company reports no signs of slowing. By the Pew research, more than half of smart phone users get health information (62%) and navigating through apps with worldwide trends up to 4.4 billion by 2017. Fortunately, this complex web of information (social media, app development, short messaging system (SMS) or text messaging, and GPS locators) can assist in continuous surveillance, information sharing, education and training, risk communication, and modeling to help predict critical nodes for quick response to potential emerging threats.
In fact, smart phones assisted disaster relief in the 2010 Haiti Earthquake by SIM cards to track population movements during cholera outbreaks. Smart phones were also used for emergency reporting systems on mosquito-borne Dengue virus to help predict epidemics and model predictors as an “early warning” device. Smart phone apps also helped educate and train first responders and clinicians. Social media activity predicted and tracked outbreaks of syndromic flu. This same network can be used to receive population surveys as well as risk communication to alleviate fear-inspired flight of misinformation. In short, leveraging this technology can enable enhanced communication, surveillance, intelligence and population management to protect health and national security.
Further, there have been recent advances in genetic sequencing, sequence analysis, and mathematical epidemiology with the convergence of several exponential trends that are cost effective and user friendly improving disease surveillance and response. This meaningful use of the bioinformatics analyses can also be performed now on web based or local systems. Additionally, DNA sequencers can diagnose disease and monitor new outbreaks. The readily available use of data collection, analyses and communication have spurred real-time access via internet, cell phone, and social media arenas. While capabilities are continually maturing, coordinated efforts to readily deploy and train these existing technologies are still in need. Specifically, Los Alamos National Laboratory (LANL) has developed tools and applications in each of these areas (genomics, disease outbreak analysis and forecasting, and the integration of genomic data into epidemiological analyses).
Empowering the Development of Genomics Expertise (EDGE) is a highly adaptable bioinformatics platform that allows laboratories to quickly analyze and interpret genomic sequence data (Figure 1). The bioinformatics platform allows users to address a wide range of use cases including assay validation and the characterization of novel biological threats within clinical and complex environmental samples. It is designed to allow users with minimal expertise in bioinformatics to be able to conduct a number of sophisticated analyses. It also aligns to real world use cases, makes use of open source (free) software tools, and runs analyses on relatively inexpensive hardware. It addresses the distinct need for advanced, yet user-friendly bioinformatics solutions, and can be provided to foreign country partners’ labs to facilitate successful adoption and use of High-Throughput Sequencing technologies. This is critical because overseas deployment of sequencers allows biological analysis closer to the acquisition point of samples increasing efficiency and effectiveness. Particularly, EDGE provides support for bio threat rapid identification, detection, and surveillance. Additionally, it can validate currently fielded polymerase chain reaction (PCR) assays as well as enable rapid design of new assays. This capability is a force health multiplier especially in Outside the Continental United States (OCONUS) and forward deployed environments where human resources, space, bandwidth, and time are limited.

Figure 1: EDGE Overview

Figure 2: EpiEarly User Interface
Currently, LANL has two concurrent projects running that are developing tools for epidemiologic analysis. A forward looking internally funded project is working to integrate data from the Los Alamos genomics group into epidemiologic models that address the extensive co-morbidities in malaria holoendemic areas of Western Kenya (Siaya). A Defense Threat Reduction Agency (DTRA) project is geared towards making the necessary diversity of epidemiologic tools available to the Analysis, Planning, Operations and Response communities through DTRA’s Biosurveillence Ecosystem. This project also addresses the uncertainty quantification that is necessary to provide decision support. Five modeling and analysis tools and one uncertainty quantification tool are being provided. The first three (EpiEarly, EpiRapid, EpiZoo) are available in a useable form currently. Two (EpiCast and EpiRapid) are in development. Because there is no “one-size fits all model” in mathematical epidemiology, these tools span the range of granularity from aggregated case-count data (EpiEarly) to event-driven agent-based models (EpiAgent). These tools are designed to look at these issues at different levels and to address a range of population sizes, mitigation detail, demographic fidelity, and computation effort as well as designed to be available to the individual analyst for force protection, operational status planning, and bio-event management and mitigation.
EpiEarly is a Bayesian estimator of reproductive number and only requires the user to input a time series of cases and the diseases’ serial generation time in order to return an estimated reproductive number with certainty (Figure 2). EpiRapid and EpiZoo are geographically compartmentalized SIR-type model (susceptible, infected, recovered) with mitigations adapted to human and animals, respectively, and allow “what-if planning and analysis scenarios” to be examined (Figures 3 & 4).

Figure 3: EpiRapid User Interface
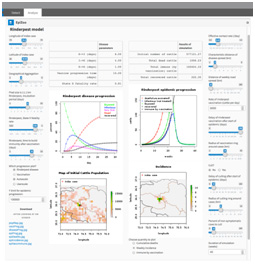
Figure 4: EpiZoo User Interface
EpiCast is an agent-based day-night model with stratified mixing groups (SIR-type), explicit agent motion, and a concept of places (home, work school, neighborhood, census tract, etc.). All motion and infection are modeled at day-night granularity. It has detailed model of disease progression to allow fine-grained simulation of interventions (e.g. vaccination, school closure, etc.). EpiAgent is fine-grained in time, space and allows behavioral simulation (mask use, social distancing), agents with definite locations (latitude/longitude type data), and infection in short-time intervals (5 min) based on the agent being in a "place." These parameters are adaptable to specific locations. EpiUpdate (in development) addresses the uncertainty within and between models providing confidence bounds that will allow analysts to appropriately qualify their work product for consideration by decision makers. This allows the data to be interpreted by the user with levels of predictability.
All these tools have interfaces that are designed to be easily understood by a user and provide sufficient flexibility that many situations of interest can be analyzed using analogy that are parallels, but without requiring that every analytic task be a research project.
Data sharing and expert analysis of incoming information are key factors to enhancing situational awareness of an unfolding event. LANL has developed an integrated suite of tools to enhance situational awareness for global disease surveillance that provide actionable information and knowledge for enhanced situational awareness during an unfolding event: Biosurveillance Resource Directory (BRD), Biosurveillance Analytics Resource Directory (BARD), Surveillance Window App (SWAP), and Epidemiologic Information Collection tool (EPIC).
The BRD is a searchable database to facilitate obtaining disease surveillance information. It contains information on disease surveillance resources worldwide. All resources are described using a comprehensive framework that describes the fundamental elements of infectious disease surveillance and contain information on how to access the data.
The BARD is a searchable database that catalogs and classifies epidemiologic model specific information to allow selection of appropriate models for use in disease prediction, monitoring or forecasting. A systematic model characterization framework has been developed to allow consistent epidemiologic model description and facilitate "apples to apples" comparison for multiple models available for the same disease.
EPIC is a data collection and visualization tool for notifiable disease data from around the world. The tool uses various scrapers to collect information from national health websites and digests it into a database that allows the data to be easily visualized, compared and exported (Figure 5).
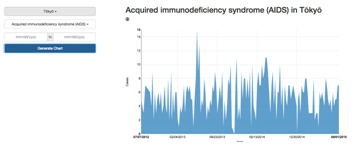
Figure 5: Epic User Interface
The SWAP is an app developed to provide context and a frame of reference for disease surveillance information about an unfolding event through matching of user input to a library of global historical disease outbreaks (Figure 6). Each historical outbreak is a “case study” and contains information on case counts, location, data sources used (from the BRD), disease models used (from the BARD), risk factors, mitigation efforts, and other epidemiologic relevant information. The SWAP is intended to provide a frame of reference for incoming data for a particular location and time during a disease outbreak so that the user is able to understand and qualify the situation to facilitate actions to prevent, stop or mitigate the event. It can also make a short term forecast for the unfolding disease situation based upon user input data.
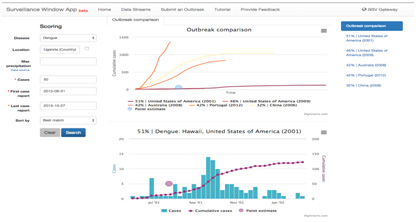
Figure 6: SWAP User Interface
All these tools described are either online, or shall be quickly available online and would require minimal modification to be able to deploy them into the hands of analysts and health officials (Table 1).
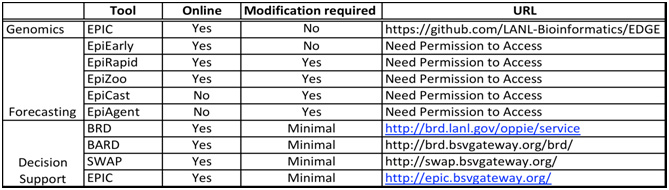
Table 1: Summary of Tools
Conclusion
In the face of budget constraints and limited resourcing, there is a need for an Integrated Global Health Surveillance and Response Program through a complex web (data sharing among veterinarians, entomologists, climatologists, microbiologists, clinicians, and public health officials) of surveillance, education, and interactive communications, otherwise known as “network of networks.” This is particularly important for expeditionary operations and Army Health Support of Operations in order to enhance Department of Defense (DoD) capabilities and to minimize population disease outbreaks that create geopolitical and socioeconomic instability. However, this concept can also affect large populations in times of war (WWI, WWII, Vietnam, Iraq, Afghanistan, and Syria) and peace (natural disasters and public health emergencies) based on strategic goals of governmental states around the world. Technologies and existing data science platforms such as those expressed by LANL provide up to date resources towards a more multi-source approach on population health.
Material has been reviewed by the Walter Reed Army Institute of Research. There is no objection to its presentation and/or publication. The opinions or assertions contained herein are the private views of the author, and are not to be construed as official, or as reflecting true views of the Department of the Army or the Department of Defense. Research was conducted in an AAALACi accredited facility in compliance with the Animal Welfare Act and other federal statutes and regulations relating to animals and experiments involving animals and adheres to principles stated in the Guide for the Care and Use of Laboratory Animals, NRC Publication, 2011 edition.
About the Author(s)
Comments
Good morning visitors! This…
Good morning visitors! This information is very helpful for me, so thanks for sharing it! I have generalized panic and anxiety disorder, and only these cool therapists have told me of many resources and useful meditations. They also taught me the process of analyzing my thoughts. During the pandemic they helped me to design a routine and to stay on track. I would recommend their https://us.calmerry.com/blog/eating-disorders/12-warning-signs-of-eating-disorders-you-shouldnt-ignore/ agency to everyone who is searching for any kind of positive support in their hard life.
